{flow} provides tools to visualize as flow diagrams the logic of functions, expressions or scripts and ease debugging.
Use cases are :
- Deciphering other people’s code
- Getting more comfortable with our own code by easing a visual understanding of its structure
- Documentation
- Debugging
- Inspect unit test results
- Providing a higher level view of an algorithm to collaborators
- Education
Installation
Install from CRAN with:
install.packages("flow")Or install development version from github:
remotes::install_github("moodymudskipper/flow")Example
Using default nomnoml engine
flow_view(rle)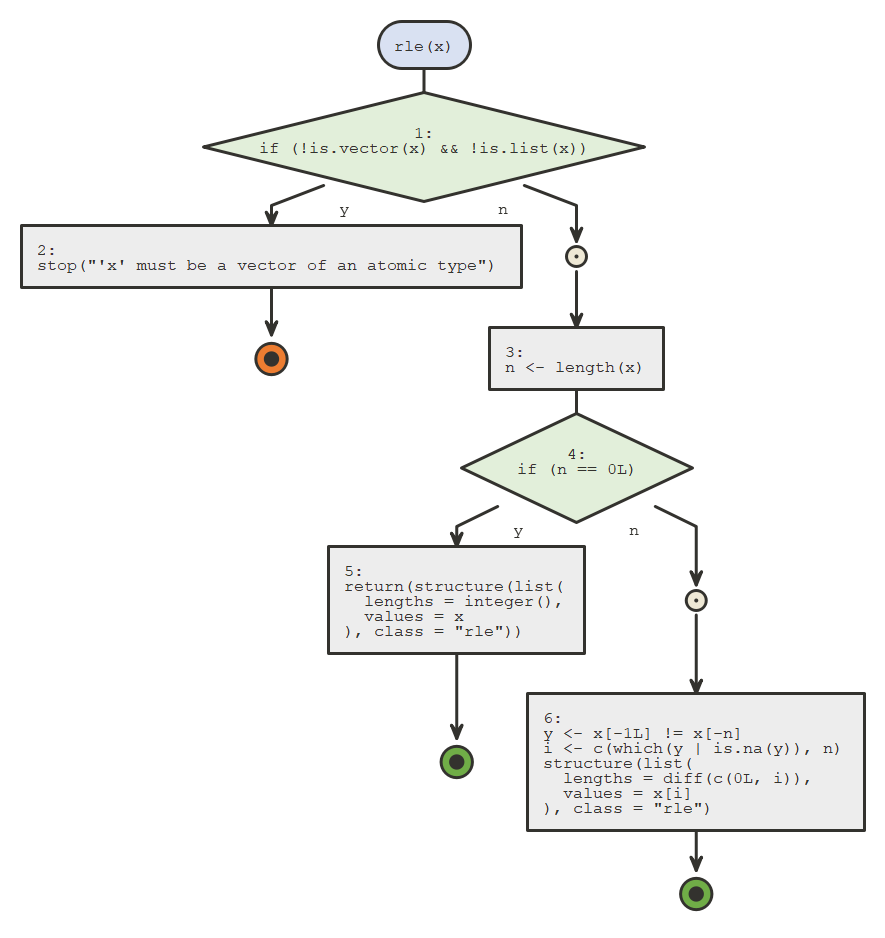
nomnoml
Using plantuml engine (make sure the {plantuml} package is installed).
flow_view(rle, engine = "plantuml")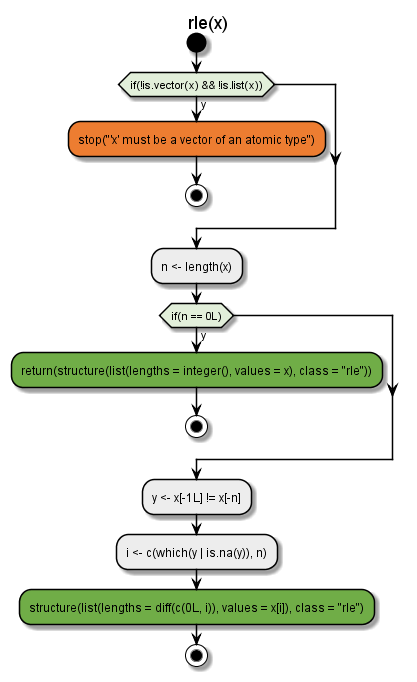
plantuml
Additional functions
-
flow_run()to display not only the diagram, but the logical path taken by a specific call -
flow_compare_runs()display the logical path of 2 calls to see where they diverge -
flow_debug()/flow_undebug()to use basically useflow_run()on a function wherever it’s called -
flow_view_vars()to display the dependencies between variables in a function -
flow_view_deps()to display recursively all the functions that your function calls -
flow_view_uses()to display recursively all the functions that call your function -
flow_view_shiny()to display the modular structure of your shiny app -
flow_view_source_calls()to display dependency tree of scripts sourcing each other -
flow_doc()to build a package’s documentation using flow diagrams -
flow_test()to show what happens in your unit tests -
flow_embed()to embed diagrams in your documentation.
See more in vignettes.
Notes
Make sure to check the vignettes for a detailed breakdown of all features.
{flow} is built on top of Javier Luraschi’s {nomnoml} package, and Rainer M Krug ’s {plantuml} package, the latter only available from github at the moment.